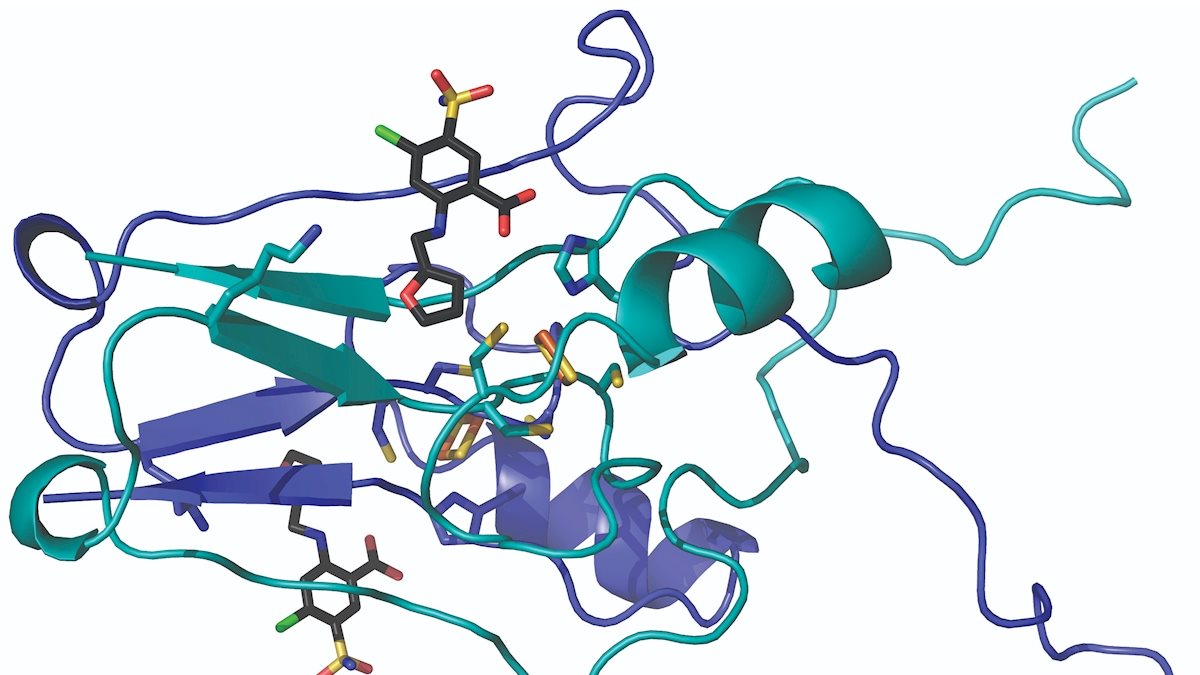
Medications attach to the proteins in our bodies the way spacecrafts dock into the International Space Station. Describing that process in detail can reveal a lot about how the medications work—and what form new medications should take.
Researchers at West Virginia University have mapped the crystal structure of a protein that resides in our cells and determined—for the first time—how a drug latches onto it. The findings appear in Communications Chemistry, a Nature research journal.
The study—funded by the West Virginia Clinical and Translational Science Institute—centered on a protein called “mitoNEET.” MitoNEET inhabits the outer membrane of our mitochondria, which act like power plants that energize our cells.
“MitoNEET is a novel therapeutic target for metabolic-based diseases and could possibly lead to disease-modifying treatments for Alzheimer’s disease and stroke,” said Werner Geldenhuys, an associate professor in the School of Pharmacy and School of Medicine. He and his colleagues—including Aaron Robart, an assistant professor in the WVU School of Medicine, John Hollander, assistant dean for professional programs in the WVU School of Medicine, and Timothy Long, an associate professor in the Marshall University School of Pharmacy—carried out the project.
“This protein has been implicated in a lot of diseases that are very tough to tackle: things like diabetes, stroke, heart disease,” Robart said. “We don’t actually know what the protein does yet, but it hangs out in proximity to the powerhouse of the cell, and all of these diseases have an energy-flow theme to them.”
To explore the role mitoNEET plays in our energy processes, the researchers isolated mitoNEET from both bacterial overexpression and animal models. Then they synthesized 11 molecules similar to furosemide—a common diuretic sold under the brand name LASIX—and exposed the mitoNEET to them.
After the molecules bonded to the mitoNEET, the researchers built atom-by-atom maps of the pairings. They remotely controlled Argonne National Laboratory’s Advanced Photon Source—which bombards samples with ultra-bright, high-energy X-rays—to reveal precisely how the molecules came together.
The team discovered that the molecules docked into a cluster of iron and sulfur atoms that made up part of the protein. Raisa Nuñez, an undergraduate participating in the Research Apprenticeship Program, collected preliminary structural data. “This highlights that significant scientific discovery can come at any career level,” Robart said.
“These findings are of importance as they allow us to continue to understand the role played by mitochondria and bioenergetics in many disease states,” Hollander said. “The modulation of mitochondrial function through targeted therapeutics may be a critical avenue of drug discovery."
Understanding mitoNEET’s cellular function could improve the performance of drugs that work by altering the protein’s activity. For example, adding an extra oxygen group to a drug’s molecular structure could dramatically tighten its bond to mitoNEET and eliminate unintended binding to other cellular proteins.
The potential upshot for patients who take the drug? Better symptom relief.
“The success of this project really illustrates how approaches that are considered basic science can provide considerable insight into clinical problems,” said Michael Schaller, who chairs the School of Medicine’s Department of Biochemistry. “It also demonstrates the power of tackling problems as teams consisting of members with very different expertise.”
Research reported in this publication was supported by the West Virginia Clinical and Translational Science Institute. WVCTSI is funded by an IDeA Clinical and Translational grant from the National Institute of General Medical Sciences, under Award Number U54GM104942, to support the mission of building clinical and translational research infrastructure and capacity to impact health disparities in West Virginia.
Citation
Title: Crystal structure of the mitochondrial protein mitoNEET bound to a benze-sulfonide ligand
